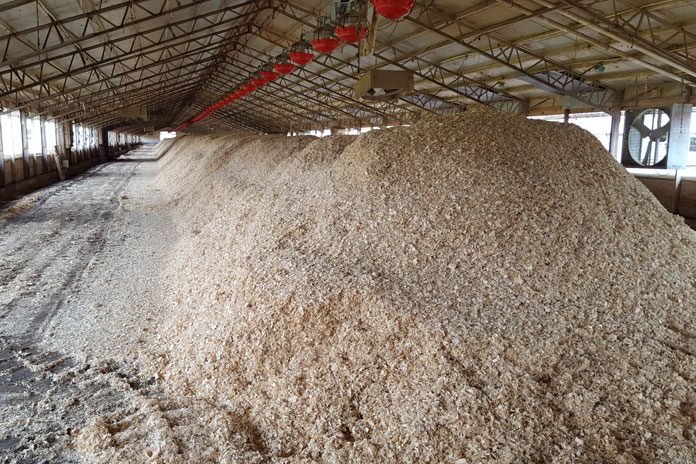
There is a well-recognised problem with both the supply of broiler litter material in some regions, and with the disposal of spent litter. This is driving a trend towards increased reuse of litter to raise successive chicken batches.
One of the risks associated with this practice is carryover of litter-borne viral and bacterial pathogens between successive batches of chickens. This risk has been confirmed and quantified for some viral pathogens, using a chick bioassay developed for this purpose. The chick bioassay has also been used successfully to determine the efficacy of litter treatments such as pasteurising by heaping or windrowing between batches of chickens in reducing virus transmission between batches in used litter. While the bioassay has proved a useful research tool, in practice it is complex and expensive to implement and it would be ideal if a direct test of viral load in litter could be developed. Quantitation of viral load using quantitative real-time PCR (qPCR) is an attractive option but to be effective requires resolving of three issues:
-Ensuring adequate representation of the original environmental material in the final nucleic acid sample assayed;
-Ensuring that the PCR reaction is not compromised by inhibitors;
-Ensuring that the reaction is measuring infectious virus, not inactivated virus with intact nucleic acids.
As part of a project to investigate these issues and determine the feasibility and utility of direct assay of viral pathogens in poultry litter using qPCR we needed first to develop suitable qPCR tests for 5 viruses and develop effective methods extraction and quantitation of viral nucleic acids from broiler litter. This paper reports our findings in this regard.
Fully quantitative Taqman® qPCR assays with plasmid-based standard curves were used to quantify Marek’s disease virus (MDV), infectious laryngotracheitis virus (ILTV), chicken anaemia virus (CAV), Fowl adenovirus (FAdV) and infectious bursal disease (IBDV). MDV ILTV and FAdV are double stranded DNA viruses, CAV is a single stranded DNA virus and IBDV is a double stranded RNA virus. Methods were either adapted from published assays or developed by the project as follows: MDV; CAV; FAdV, modified HEX-S; ILTV, IBDV developed by project from GenBank sequence data of the VP2 gene.
Specific pathogen free (SPF) litters (LT11-C-LP1) based on pine shavings, hardwood shavings and rice hulls were generated by placing SPF chickens on these litters for 42 days then storing at -20°C and following drying for 7 days at 37°C until required. Infective litter was generated by raising broilers vaccinated with MDV, ILTV, IBDV, CAV and FAdV live vaccines on pine shavings for 42 days (LT11-C-LP2). Stored infective pine shavings from the study of were also used (Samples LT09-C-CB9).
Study 1 comprised a number of small-scale tests to demonstrate isolation of a DNA (MDV) and an RNA virus (IBDV) from naturally contaminated litter and testing of various preparation steps prior to nucleic acid extraction. These treatments included washing the litter with MilliQ water, or TE buffer containing 0.15% Tween-80 to release the virus from the litter (shaking or blending); washing the litter with 10% beef extract then precipitation of virus with 8% PEG 6000.
Study 2 comprised a formal factorial experiment testing the effects of litter washing method (TE buffer plus 0.15% Tween-80 or 10% beef extract), wash times (2 hr and 16 hr at 4°C), blending (blend, no blend) and bead beating (bead beat, no bead beat) on detection of MDV and IBDV in DNA/RNA extracted from unstrained slurry, or the retentate or filtrate following straining of the slurry.
Study 3 tested for the presence of PCR inhibitors in litter of the different types by addition of fixed amounts of viral DNA to serial dilutions of extracted litter material and then investigated the effects of polyvinylpolypyrrolidone (PVPP) in overcoming inhibition.
Study 4 was a large factorial experiment in which fixed amounts of commercially available vaccine for 4 of the viruses and infective dust for MDV were added to 100 g of SPF litters of different types and recovery in different fractions determined by qPCR following a range of preparatory steps. Samples of the pine shavings, hardwood shavings and rice hulls litters were incubated in TE buffer plus 0. 15% Tween-80 on ice with shaking for two hours (blending twice during this time) then filtered through a fine nylon mesh (approximately 1mmq). The retentate (R) was retained and the filtrate centrifuged at 17,500g for 30mins at 4C, to provide a supernatant (S) and pellet (P). The R, S and P fractions were then subjected to the following treatments in a complete factorial design: Bead beating (BB) for 5 minutes or not; and treatment with PVPP or not.Total nucleic acids were extracted using magnetic bead technology (MagMaxTM Total RNA isolation kit Ambion) manually or in the automated Kingfisher Flex 96 (Thermo Fischer Scientific) although the Qiagen QIAmp DNA stool extraction kit was also tested in Study 1. The final eluate was diluted 1:10 and used as the template in the qPCR reaction for all 5 viral assays. Assay results were generally expressed as viral copies (VC) per unit.Where relevant the significance of fixed effects and their interactions in the various studies were determined by ANOVA with means separation by protected Student’s T test, or in the case of ratios, by contingency table analysis. Analyses were performed with JMP 9. Means are presented ± SEM.
Results
Study 1: As this comprised a series of small tests, formal statistical analysis was not attempted. Nucleic acid targets of both MDV (DNA virus) and IBDV (RNA virus) were successfully recovered from pine shaving litter material naturally contaminated by infected chickens. Significant amounts of both were detected in the retentate, filtrate and pellet fractions following litter preparation. Use of Tween-80 to wash the litter provided better overall results for both viruses than water or the beef extract. Water failed to release IBDV from the litter sample with the majority of the viral RNA detected in the initial litter strainings, however it was successful in releasing MDV. Conversely IBDV was satisfactorily detected using beef extract whilst MDV was not. Blending and shaking samples in Tween 80 produced high recoveries for both viral targets so this was explored further in Study 2.
Study 2: MDV, but not IBDV target sequence amplified successfully in this experiment. For MDV the overall mean value for viral load was 8417 ± 1813 VC/g litter for all preparation methods and fractions. Increasing wash time from 2 hr to 16 hr at 4°C significantly reduced the number of positive samples from 96% to 67% (P=0.006). Washing with Tween buffer resulted in significantly higher MDV recovery than washing in beef extract buffer (11,585 v 5,245 VCN/g litter, P=0.05). More viral DNA was recovered in the filtrate (15,161 VC/g litter) than the solid retentate after straining (1,275 VC/g litter) (P=0.004) with unstrained slurry intermediate (8,815 VC/g litter). There were no other significant main effects although there were trends towards improved recovery with blending, and reduced recovery with bead beating.
Study 3: Initial spiking tests using a 7 x 10-fold serial dilution of DNA extracted from pine shavings, rice hulls and hardwood shavings indicated that the qPCR reaction for MDV was inhibited in undiluted samples for all litter types, with amplification of the target first occurring at dilutions of 1:10 for pine shavings and rice hulls and 1:1000 in hardwood shavings. When DNA was extracted from infective litter dried at 37 ̊C rather than stored at –20°C, similar results were obtained except that amplification occurred at a dilution of 1:100 in the hardwood shavings. The spike test was repeated using a 6 x 2-fold serial dilution of DNA extracted from hardwood shavings with 6.6% w/v PVPP added to the extracted DNA sample and mixed for 5 minutes, 1 hour or overnight at room temperature. Amplification of the target product occurred at dilutions of 1:4, 1:2 and neat respectively. In all of the tests, once amplification occurred, there was no effect of further dilution on the amount of target recovered when adjusted for the dilution factor.
Study 4: The dataset comprised 144 qPCR results. CAV, IBDV, ILTV, and MDV, but not FAdV were readily detected in all three litter types with overall percentages of positive qPCR results of 69%, 43%, 22% and 26% respectively (P<0.001). When no PVPP was used the percentage of positive samples was significantly lower from hardwood (6%) than rice hulls (50%) or softwood, (47%) (P<0.0001) but this effect was completely removed by inclusion of PVPP with values of 48%, 48% and 42% respectively, indicating a selective effect of PVPP in overcoming the PCR inhibition observed with hardwood shavings. The effect of PVPP was evident for all viruses but was greatest for CAV and least for IBDV. For the three DNA viruses no amplification occurred at all on hardwood shavings in the absence of PVPP. The pellet had the highest ratio of positive samples (47%) followed by the supernatant (41%), then the retentate (33%) (P=0.07). PVPP increased the percentage of positive samples in the pellet (P=0.008). Bead beating increased the proportion of positive samples from 35% to 45% (P=0.09). This was observed for all viruses except ILTV for which there was no difference with or without bead beating. The mean measured recovery rates of added virus were 18.0, 10.4, 0.065 and 0.052 % for IBDV, CAV, ILTV and MDV respectively (P<0.001). The virus recovery rate was not significantly affected by any of the other factors in the experiment.
This work confirms other published work demonstrating that viral nucleic acids can be reliably amplified from poultry litter. Clearly the type of litter is important, with hardwood based litter material containing PCR inhibitors. Fortunately these were able to be neutralised with PVPP. For all viruses significant amounts were found in the retentate after filtration, and in both the pellet and supernatant fractions following centrifugation of the filtrate. The results indicate that a method based on washing samples with TE containing 0.15% Tween-80 followed by bead beating and PVPP treatment would enable detection of both DNA and RNA viruses from litter with the greatest concentration of virus found in the pellet fraction after centrifugation. Detectable virus recovery was good for IBDV and CAV, but low for the dsDNA viruses ILTV and MDV. The third dsDNA virus, FAdV, was unable to be recovered at all. Work is ongoing to resolve the low recovery rate of dsDNA viruses and to simplify the litter processing and DNA extraction further.